应用介绍
BindCraft 是一个用于蛋白质设计、分子对接和结构建模的开源软件工具,旨在通过优化蛋白质-配体相互作用、提高设计的准确性和效果,辅助科研人员在药物设计、蛋白质工程等领域开展高效的计算工作。该工具主要应用于蛋白质-小分子或蛋白质-蛋白质的相互作用建模,以及通过机器学习和其他计算方法来设计和优化蛋白质结构。
加载环境
export PATH=/opt/app/singularity/bin:$PATH
使用说明
实例演示
- 申请计算节点
salloc --gres=gpu:1
ssh XXX
- 创建个人目录,拷贝软件目录
mkdir -p $HOME/user_test
cp -r /opt/app/bindcraft $HOME/user_test/bindcraft
chmod 777 -R $HOME/user_test/
- 启动 singularity 容器***
singularity run --nv -B $HOME/user_test/:/content -B $HOME/user_test/bindcraft:/content/bindcraft -B /opt/app/nvidia/550.54.14:/usr/local/nvidia /opt/app/sif/bindcraft.sif
- 加载容器内 conda env 环境
source /opt/conda/bin/activate
conda activate BindCraft
- 执行 demo 测试
python -u /content/bindcraft/bindcraft.py --settings '/content/bindcraft/settings_target/PDL1.json' --filters '/content/bindcraft/settings_filters/default_filters.json' --advanced '/content/bindcraft/settings_advanced/default_4stage_multimer.json'
常见问题
Q1:
ARNING: 'model_1_multimer_v3' not found
WARNING: 'model_2_multimer_v3' not found
WARNING: 'model_3_multimer_v3' not found
WARNING: 'model_4_multimer_v3' not found
WARNING: 'model_5_multimer_v3' not found
......
tage 1: Test Logits
Traceback (most recent call last):
File "/iridisfs/scratch/binder/BindCraft/./bindcraft.py", line 108, in <module>
trajectory = binder_hallucination(design_name, target_settings["starting_pdb"], target_settings["chains"],
File "/iridisfs/scratch/binder/BindCraft/functions/colabdesign_utils.py", line 97, in binder_hallucination
af_model.design_logits(iters=50, e_soft=0.9, models=design_models, num_models=1, sample_models=advanced_settings["sam
ple_models"], save_best=True)
File "/home/.conda/envs/BindCraft/lib/python3.9/site-packages/colabdesign/af/design.py", line 346, in design_lo
gits
self.design(iters, **kwargs)
File "/home/.conda/envs/BindCraft/lib/python3.9/site-packages/colabdesign/af/design.py", line 340, in design
self.step(lr_scale=lr_scale, num_recycles=num_recycles,
File "/home/.conda/envs/BindCraft/lib/python3.9/site-packages/colabdesign/af/design.py", line 206, in step
self.run(num_recycles=num_recycles, num_models=num_models, sample_models=sample_models,
File "/home/.conda/envs/BindCraft/lib/python3.9/site-packages/colabdesign/af/design.py", line 89, in run
model_nums = self._get_model_nums(num_models, sample_models, models)
File "/home/.conda/envs/BindCraft/lib/python3.9/site-packages/colabdesign/af/design.py", line 71, in _get_model_nums
ns = [ns[n if isinstance(n,int) else ns_name.index(n)] for n in models]
File "/home/.conda/envs/BindCraft/lib/python3.9/site-packages/colabdesign/af/design.py", line 71, in <listcomp>
ns = [ns[n if isinstance(n,int) else ns_name.index(n)] for n in models]
IndexError: list index out of range
A1: FIX
## 缺少 alphafold 权重
https://storage.googleapis.com/alphafold/alphafold_params_2022-12-06.tar
cp to " ~/bindcraft/params"
Q2:
ERROR: Cannot open file "/content/drive/My Drive/Bindcraft/PDLl/Trajectory/PDL1_l141_s14957.pdb""
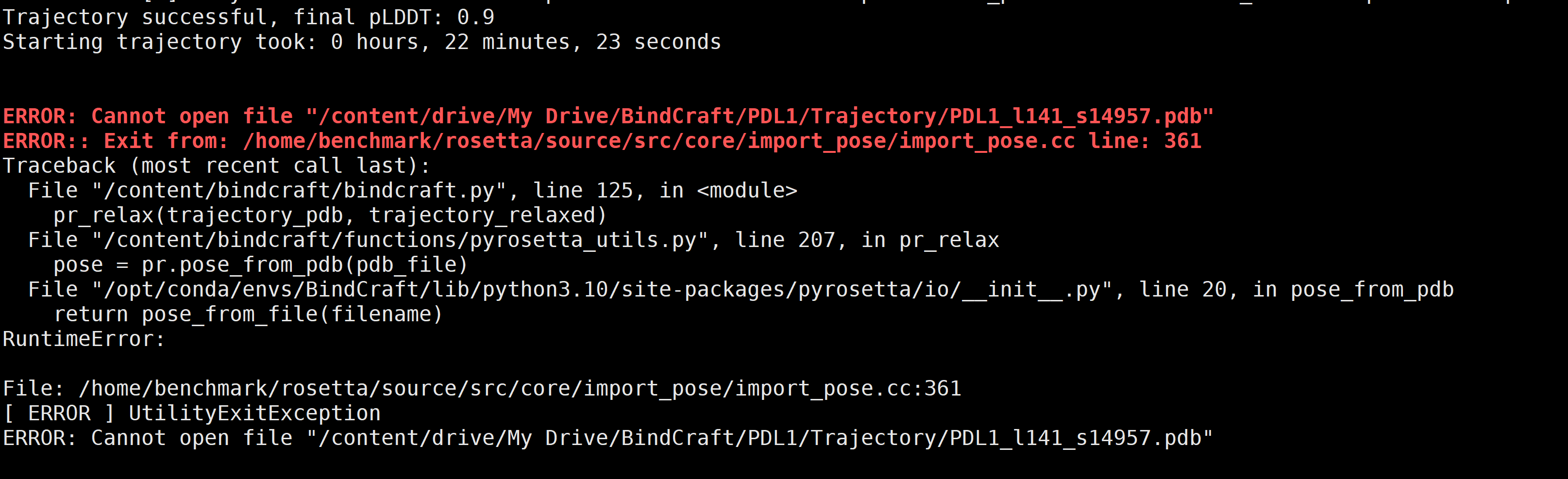
A2: FIX
## 路径中不能有空格
~/bindcraft/settings_target/PDL1.json
"design_path": "/content/drive/MyDrive/BindCraft/PDL1/"
